Principal Component Analysis (PCA) is a dimensionality reduction technique commonly used in data analysis and machine learning. One of its primary objectives is to capture the most variance in the data while reducing the dimensionality of the dataset. Variance is a statistical measure that quantifies the spread or dispersion of data points. In the context of PCA, capturing the most variance means retaining the most important information in the dataset while reducing its complexity.
PCA transforms the original features of a dataset into a new set of orthogonal (uncorrelated) variables called principal components. These principal components are linear combinations of the original features. This means that each principal component is created by taking a weighted sum of the original features.
The amount of variance captured by each principal component is quantified by its eigenvalue. PCA orders these principal components in such a way that the first principal component captures the most variance, the second captures the second most, and so on. The first principal component has the highest eigenvalue. Subsequent principal components have decreasing eigenvalues, indicating their decreasing importance in explaining the variance.
By selecting a subset of the principal components that capture the most variance, we can maintain the essential information present in the original data. This is particularly useful for tasks like data visualization, noise reduction, and feature engineering in machine learning.
In summary, PCA captures the most variance in the data by identifying and prioritizing the principal components that explain the largest amount of variation in the dataset. This allows us to reduce the dimensionality of the data while retaining the most important information.
Assumptions
PCA is based on the Pearson correlation coefficient framework and inherits similar assumptions. The Pearson correlation coefficient, often denoted as “r” or “Pearson’s r,” is a statistical measure used to quantify the strength and direction of the linear relationship between two continuous variables. It assesses how well the relationship between these variables can be described by a straight line.
- Sample size: Minimum of 150 observations and ideally a 5:1 ratio of observation to features.
- Correlations: The feature set is correlated, so the reduced feature set effectively represents the original data space.
- Linearity: All variables exhibit a constant multivariate normal relationship, and principal components are a linear combination of the original features.
- Outliers: No significant outliers in the data as these can have a disproportionate influence on the results.
- Large variance implies more structure: high variance axes are treated as principal components, while low variance axes are treated as noise and discarded.
Step-by-step Procedure for PCA:
- Standardize the Data:
- The first step in PCA is to standardize your dataset to have a mean of 0 and a standard deviation of 1. This ensures that each feature contributes equally to the computation of principal components. Formula: zi= (xi-μ)/σ Where:
- zi = standardized data
- xi = original data point
- μ = mean of the feature
- σ = standard deviation of the feature
- The first step in PCA is to standardize your dataset to have a mean of 0 and a standard deviation of 1. This ensures that each feature contributes equally to the computation of principal components. Formula: zi= (xi-μ)/σ Where:
- Calculate the Covariance Matrix:
- The covariance matrix captures the joint variability of your data. If your dataset is
m x n(m samples and n features), the resulting matrix will ben x n.
- The covariance matrix captures the joint variability of your data. If your dataset is
- Calculate the Eigenvectors and Eigenvalues:
- Eigenvalues represent the magnitude (or variance) captured by each principal component.
- Eigenvectors represent the direction of each principal component in the original feature space.
- For PCA, you’ll compute the eigenvectors and eigenvalues of the covariance matrix.
- Sort Eigenvectors by Eigenvalues in Descending Order:
- This step is crucial because the eigenvector with the highest eigenvalue is the principal component that captures the most variance in the data.
- Choose the Number of Principal Components:
- Based on the problem requirements and the explained variance, decide how many principal components you want to retain. A common approach is to compute the cumulative explained variance and pick a number of components that explain, say, 95% of the total variance.
- Project the Original Data:
- Multiply the standardized data by the top k eigenvectors (where k is the number of principal components you decided to retain). This will give you a new dataset with reduced dimensions, where each row is a linear combination of the original features weighted by the principal components.
- Use the Transformed Data:
- You can now use the reduced-dimensional dataset for various tasks like clustering, regression, classification, or visualization.
Mathematics behind PCA
The whole process of obtaining principle components from a raw dataset can be simplified into six parts :
- Take the whole dataset consisting of d+1 dimensions and ignore the labels such that our new dataset becomes d-dimensional.
- Compute the mean for every dimension of the whole dataset.
- Compute the covariance matrix of the whole dataset.
- Compute eigenvectors and the corresponding eigenvalues.
- Sort the eigenvectors by decreasing eigenvalues and choose k eigenvectors with the largest eigenvalues to form a d × k dimensional matrix W.
- Use this d × k eigenvector matrix to transform the samples onto the new subspace.
In a nutshell, to transform the features into Principal Components, we multiply the features with the eigenvectors element by element. Note that each of the principal components contains information from all the features. Transform Principal Components back into features, multiply the transpose of the original data set by the transpose of the feature vector. The feature vector is simply a matrix that has as columns the eigenvectors of the components that we decide to keep.
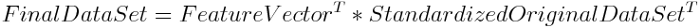
[A covariance matrix is a square matrix that summarizes the covariances between multiple variables. Covariance is a measure of how two random variables change together. The covariance between two variables (X and Y) measures whether, when X increases, Y tends to increase or decrease.
An eigenvector is a vector whose direction remains unchanged when a linear transformation is applied to it.]

